G-protein coupled receptors (GPCRs) are the largest superfamily of membrane proteins in the human genome and a significant target for more than 30% of pharmaceutical drugs. Here we use molecular docking and molecular dynamics simulations to explore the determinants responsible for α/γ-peptide binding selectivity of the Y4R receptor.
Motivation and problem definition
The neuropeptide Y (NPY) receptor family comprises four physiologically relevant class A GPCRs, designated Y1R, Y2R, Y4R, and Y4R. The endogenous ligands of NPY receptors are the homologous linear peptides neuropeptide Y (NPY), peptide YY (PYY), and pancreatic polypeptide (PP), all consisting of 36 amino acids. Because of its role in appetite suppression, the Y4R receptor is a very attractive target for the design of new therapeutic compounds for fighting obesity. Unlike small molecules, peptides exhibit high conformational flexibility in their unbound states due to the large number of rotatable bonds along the backbone and in the side chains. In contrast, the receptor binding pocket imposes a stringent constraint on the conformation of these peptides. To date, relatively few studies that allow us to gain insight into the molecular basis of ligand recognition on Y4R-peptide systems exist.
Computational methods such as molecular docking and molecular dynamics simulations are essential tools for investigating protein-ligand interactions and subsequent characterization of binding pockets. Providing details at an atomistic level of the main features related to the binding process will facilitate the rational development of Y4R selective ligands.
Methods and codes
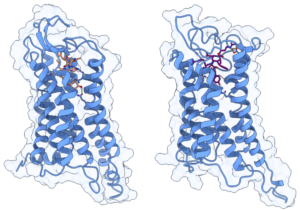
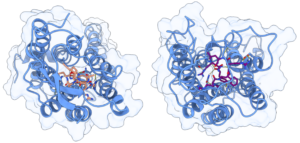
To test the structural stability of the Y4R-peptide complex, we performed 2 µs of unconstrained molecular dynamics simulations conducted in triplicate for two α/γ-peptide enantiomers with different partial agonist binding affinity and potency. The topology for the receptor was generated using the AMBER99SB-ILDN force field and peptides were parameterized using the generalized AMBER force field (GAFF). Running on one A40 GPU with Gromacs2021.5, an average simulation speed of more than 130 ns/day was obtained for GPCR systems comprising about 145,000 atoms.
Results
During the MD simulations, different electrostatic interactions between the positively charged arginine residues in the peptides and polar residues in the Y4R receptor, were detected. The most frequent interactions include polar contacts with residues T2.61, Y2.64, and Q3.32, hydrogen bonds with L2.60 and S3.28, while cation-π interactions occurred with Y2.64. The occurrence of a salt bridge involving the residue R33 and residue E6.58 in Y4R was observed for the α/γ-peptide presenting higher binding affinity and potency. These insights into Y4R ligand recognition can enable structure-based drug discovery that targets NPY receptors.
Outreach
A publication “Exploring α/γ-peptides containing phenylcyclobutyl-γ-residues for selective binding and activation of the neuropeptide Y Y4 Receptor” is in preparation.
The project was supported by the German Science Foundation (DFG) within the Research Training Group 1910 – Medicinal Chemistry of Selective GPRC Ligands.
Researcher’s Bio and Affiliation
Jacqueline Calderón studied chemistry and chemical engineering at UCV in Venezuela. She joined FAU as a PhD student in January 2020 and works in the group of Prof. Dr. Tim Clark at the Computer Chemistry Center.
